Hyper Parameter Tuning#
1. Key Hyperparameters in t-SNE#
Hyperparameter |
Role |
Typical Range |
Effect |
|---|---|---|---|
|
Controls the effective neighborhood size |
5–50 |
Low → focuses on very local structure; High → includes more global structure |
|
Step size for gradient descent |
10–1000 |
Too low → slow convergence; too high → unstable embedding |
|
Number of iterations |
250–1000+ |
Too few → embedding may not converge |
|
Initial embedding |
‘random’ or ‘pca’ |
PCA init often gives faster convergence and more stable results |
|
Distance metric |
‘euclidean’, ‘cosine’, etc. |
Should reflect similarity in high-D space |
|
Temporarily inflates \(p_{ij}\) |
Default 12 |
Helps separate clusters in early optimization |
2. How to Tune t-SNE#
a) Perplexity#
Most important hyperparameter.
Guideline:
Smaller datasets → lower perplexity (5–30)
Larger datasets → higher perplexity (30–50)
Too low → clusters may fragment.
Too high → clusters may merge or lose local detail.
b) Learning Rate#
Controls the speed of optimization.
Start with default (~200) and adjust if:
Points collapse into a tight blob → increase learning rate.
Points scatter randomly → decrease learning rate.
c) Number of Iterations (n_iter)#
t-SNE needs enough iterations for gradient descent to converge.
Typical: 1000–2000 iterations for 2D embedding.
Visual check: if clusters are still moving after final iteration → increase
n_iter.
d) Initialization#
Random → may give slightly different embeddings on each run.
PCA → more stable and faster convergence.
3. Practical Tips#
Try multiple perplexities: Compare embeddings to see which best preserves cluster structure.
Visual inspection is key: There is no quantitative accuracy metric.
Use PCA preprocessing: Reduce to 30–50 dimensions before t-SNE for noisy/high-dimensional data.
Fix random seed: Ensures reproducibility.
Avoid extremely large datasets: t-SNE scales poorly; consider subsampling or FIt-SNE for large data.
Intuition
Perplexity → “how many neighbors matter”
Learning rate → “how fast the points move to match neighbors”
Iterations → “how long we allow points to settle into a stable map”
Initialization → “starting configuration for optimization”
Think of t-SNE like folding a high-dimensional map onto 2D: these hyperparameters control the scale, speed, and smoothness of the folding.
import numpy as np
import matplotlib.pyplot as plt
from sklearn.datasets import make_blobs
from sklearn.manifold import TSNE
# Step 1: Generate synthetic 5D dataset with 3 clusters
X, y = make_blobs(n_samples=300, centers=3, n_features=5, cluster_std=1.0, random_state=42)
# Step 2: Define different perplexities
perplexities = [5, 30, 50]
tsne_embeddings = []
# Step 3: Apply t-SNE with different perplexities
for p in perplexities:
tsne = TSNE(n_components=2, perplexity=p, random_state=42)
X_embedded = tsne.fit_transform(X)
tsne_embeddings.append(X_embedded)
# Step 4: Plot embeddings
plt.figure(figsize=(18, 5))
for i, p in enumerate(perplexities):
plt.subplot(1, 3, i+1)
plt.scatter(tsne_embeddings[i][:, 0], tsne_embeddings[i][:, 1], c=y, cmap='viridis', alpha=0.7)
plt.title(f"t-SNE Embedding (Perplexity={p})")
plt.xlabel("Dimension 1")
plt.ylabel("Dimension 2")
plt.grid(True)
plt.show()
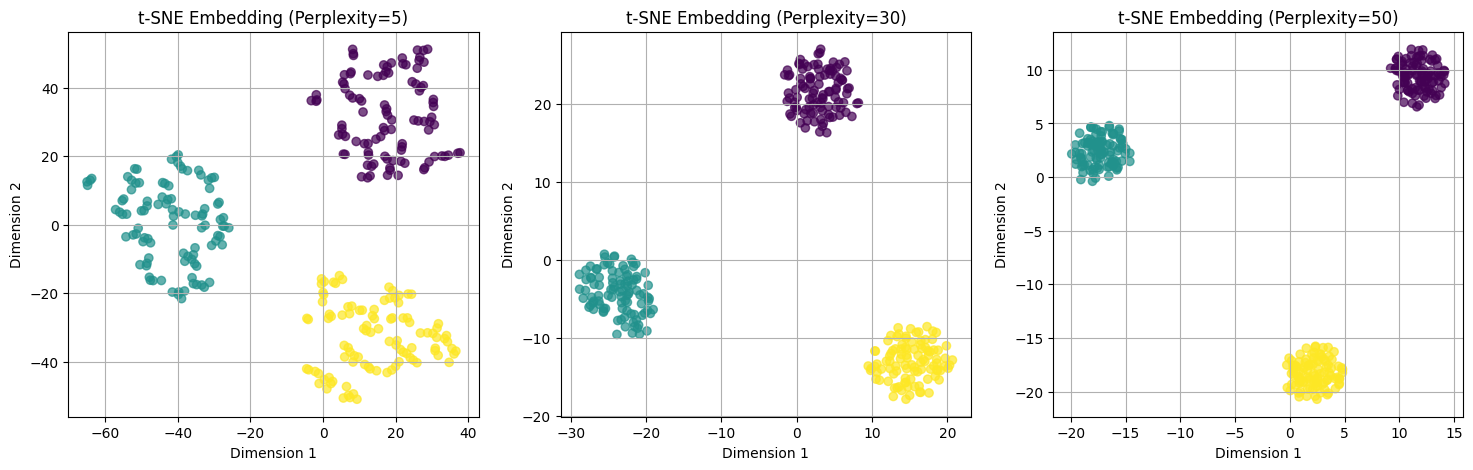
What You Will Observe#
Perplexity = 5 (small)
Very local neighborhoods are emphasized.
Clusters may appear fragmented.
Perplexity = 30 (moderate, default)
Balanced local and global structure.
Clusters are well-separated and visually meaningful.
Perplexity = 50 (large)
Neighborhoods are larger → points from different clusters may appear closer.
Global structure becomes more prominent, but local cluster details may blur.
Intuition#
Perplexity is like “how many neighbors each point cares about”.
Smaller → emphasizes fine-grained local structure.
Larger → smooths out local details, emphasizes global structure.

