Assumptions#
1. Local Neighborhood Preservation#
Assumption: Points that are close in high-dimensional space should also be close in low-dimensional space.
Implication: t-SNE is designed to preserve local structure, not global distances.
If your data doesn’t have meaningful local clusters, t-SNE may produce misleading embeddings.
2. Smooth Manifold Structure#
Assumption: High-dimensional data lies on or near a smooth, low-dimensional manifold.
t-SNE tries to “unfold” this manifold in 2D or 3D.
If the data is completely random or lacks structure, t-SNE embeddings will appear noisy.
3. Pairwise Similarity Meaningful#
Assumption: The Euclidean distance (or chosen metric) in the high-dimensional space reflects meaningful similarity.
t-SNE converts distances into probabilities, so if your distance metric is poor, t-SNE will produce misleading results.
4. Noisy Distances Handled by Heavy-Tailed Distribution#
t-SNE assumes that small distances are meaningful, but larger distances may not be reliable.
That’s why it uses a Student-t distribution in low-dimensional space:
Heavy tails allow distant points to be “pushed away” without distorting local neighborhoods.
5. Perplexity Encodes Effective Neighborhood Size#
Assumption: There exists a meaningful “local neighborhood size” in the data.
The
perplexityparameter sets this expected neighborhood size.If the chosen perplexity is too small or too large relative to actual structure, embeddings may fail to reflect true clusters.
6. Non-Determinism Assumption#
t-SNE assumes stochastic optimization is acceptable:
Different runs may produce different embeddings.
Multiple runs can reveal the robustness of clusters.
Summary Table#
Assumption |
Explanation |
|---|---|
Local structure matters |
Nearby points should stay close in embedding |
Data lies on a low-D manifold |
t-SNE tries to unfold it in 2D/3D |
Distances are meaningful |
High-D metric reflects similarity |
Noise tolerance |
Heavy-tailed t-distribution separates distant points |
Neighborhood size exists |
Controlled via |
Stochastic optimization |
Non-deterministic embedding is acceptable |
Key Intuition#
t-SNE is all about preserving local neighborhoods.
If your data has meaningful clusters locally → t-SNE will show clear clusters.
If the data is noisy or distances are meaningless → t-SNE can produce confusing embeddings.
import numpy as np
import matplotlib.pyplot as plt
from sklearn.datasets import make_blobs
from sklearn.manifold import TSNE
# Generate synthetic 3-cluster dataset
np.random.seed(42)
X, y = make_blobs(n_samples=300, centers=3, n_features=5, cluster_std=1.0)
# Apply t-SNE with different perplexities
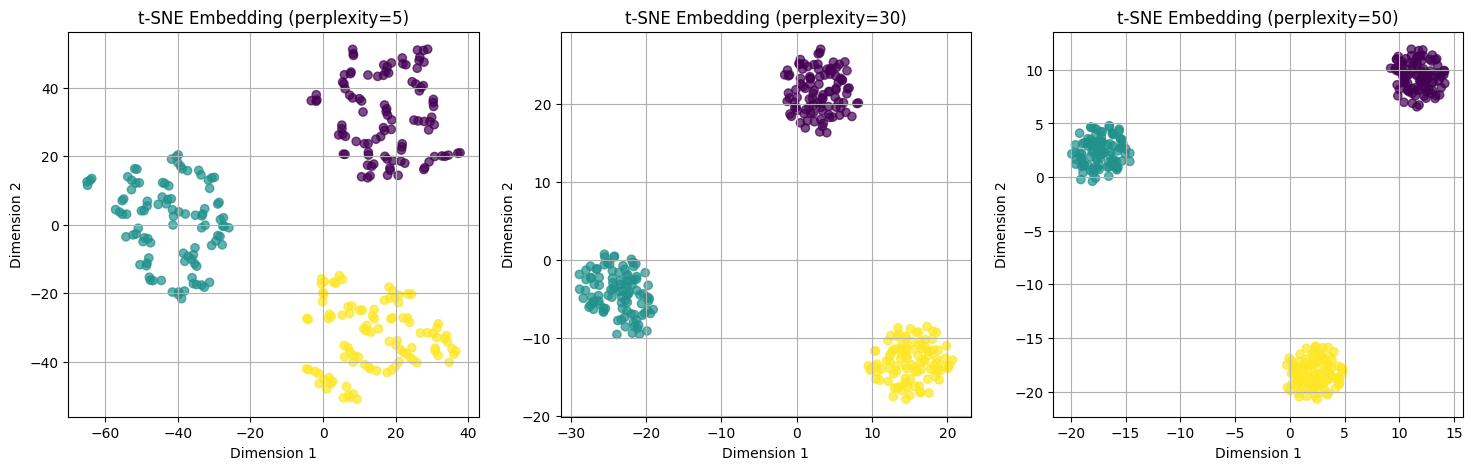
perplexities = [5, 30, 50]
tsne_embeddings = []
for p in perplexities:
tsne = TSNE(n_components=2, perplexity=p, random_state=42)
X_embedded = tsne.fit_transform(X)
tsne_embeddings.append(X_embedded)
# Plot t-SNE embeddings
plt.figure(figsize=(18, 5))
for i, p in enumerate(perplexities):
plt.subplot(1, 3, i+1)
plt.scatter(tsne_embeddings[i][:, 0], tsne_embeddings[i][:, 1], c=y, cmap='viridis', alpha=0.7)
plt.title(f"t-SNE Embedding (perplexity={p})")
plt.xlabel("Dimension 1")
plt.ylabel("Dimension 2")
plt.grid(True)
plt.show()