Local Outlier Factor#
Local Outlier Factor (LOF) is an unsupervised anomaly detection technique.
Focuses on detecting local outliers — points that are unusual relative to their neighbors.
Distinguishes between:
Local outlier: Close to a cluster but slightly different from neighbors.
Global outlier: Far from all clusters, clearly isolated.
2. Intuition Behind LOF#
LOF measures local density deviation:
Points in sparse regions compared to their neighbors → anomalies.
Uses k-nearest neighbors (kNN) to estimate local density.
Key idea: Compare a point’s density with the density of its neighbors.
If density is much lower → point is a local outlier.
3. How LOF Works#
Choose k, the number of nearest neighbors.
For each point:
Find its k nearest neighbors.
Compute the average distance to neighbors → estimates local density.
Compare local density of the point with densities of neighbors:
Low density relative to neighbors → high LOF score → outlier.
High density relative to neighbors → normal point.
LOF score interpretation:
Score ≈ 1 → normal
Score > 1 → outlier
Higher score → stronger anomaly
4. Advantages#
Detects local anomalies that global methods (like Isolation Forest) might miss.
Handles datasets with clusters of varying densities.
Based on distance metrics: Euclidean, Manhattan, Minkowski, etc.
5. Practical Usage#
Available in
sklearn.neighbors.LocalOutlierFactor.Parameters:
n_neighbors: Number of neighbors to use for density estimation.metric: Distance metric (e.g., Euclidean, Manhattan).contamination: Expected proportion of outliers.
Output:
LOF score for each sample.
Can identify local anomalies for further analysis.
Summary
LOF is useful for detecting local deviations in density.
Especially effective when data has clusters with different densities.
Uses k-nearest neighbor distances to compute local anomaly scores.
import numpy as np
import matplotlib.pyplot as plt
from sklearn.neighbors import LocalOutlierFactor
# Generate sample data
np.random.seed(42)
# Two clusters
cluster1 = np.random.randn(100, 2) + [2, 2]
cluster2 = np.random.randn(100, 2) + [7, 7]
# Outliers
outliers = np.array([[0, 0], [8, 0], [3, 9]])
# Combine data
X = np.vstack([cluster1, cluster2, outliers])
# Fit Local Outlier Factor
lof = LocalOutlierFactor(n_neighbors=20, contamination=0.05) # 5% expected outliers
y_pred = lof.fit_predict(X) # -1 = outlier, 1 = inlier
lof_scores = -lof.negative_outlier_factor_ # higher score = more anomalous
# Plot results
plt.figure(figsize=(8,6))
plt.scatter(X[:,0], X[:,1], c=['red' if i==-1 else 'blue' for i in y_pred], s=50, edgecolor='k')
plt.xlabel("Feature 1")
plt.ylabel("Feature 2")
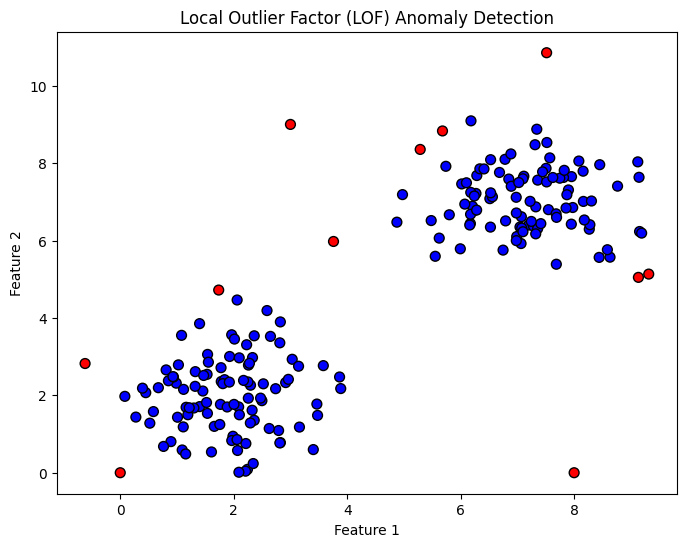
plt.title("Local Outlier Factor (LOF) Anomaly Detection")
plt.show()
# Print outlier points
print("Detected outliers:")
print(X[y_pred==-1])
Detected outliers:
[[-0.6197451 2.8219025 ]
[ 1.73534317 4.72016917]
[ 7.51504769 10.85273149]
[ 9.31465857 5.13273481]
[ 3.75873266 5.97561236]
[ 9.13303337 5.0479122 ]
[ 5.67976679 8.83145877]
[ 5.28686547 8.35387237]
[ 0. 0. ]
[ 8. 0. ]
[ 3. 9. ]]
Explanation
We generate two clusters and a few outliers.
LocalOutlierFactorcomputes LOF score for each sample based on k-nearest neighbors.Points with low local density compared to neighbors are marked as outliers (
-1).We visualize:
Blue points → normal/inliers
Red points → detected outliers
Key Notes
n_neighbors: Controls how “local” the density comparison is.Small
k→ sensitive to local anomaliesLarge
k→ more like global anomalies
contamination: Expected fraction of anomalies, helps define threshold.LOF detects local anomalies that might not be far from other clusters but are sparse relative to their neighbors.

