Workflows#
1. Input#
Dataset \(D\).
Parameters:
\(\epsilon\): neighborhood radius.
minPts: minimum number of points to form dense region.
2. Initialization#
Mark all points as unvisited.
Cluster label counter = 0.
3. Iterative process#
For each unvisited point \(p\):
Mark \(p\) as visited.
Compute neighborhood \(N_\epsilon(p)\).
If \(|N_\epsilon(p)| < \text{minPts}\): mark \(p\) as noise (temporarily).
Else:
Start a new cluster.
Increment cluster counter.
Add \(p\) and all its neighbors to this cluster.
For each point \(q\) in \(N_\epsilon(p)\):
If \(q\) is unvisited:
Mark it visited.
If \(|N_\epsilon(q)| \geq \text{minPts}\), expand cluster by merging neighbors.
If \(q\) is not yet assigned to a cluster, assign it.
4. Expansion#
Repeat neighborhood exploration until no new points can be added.
This ensures maximal density-reachability closure.
5. Completion#
Continue scanning through dataset until all points are visited.
Output:
Clusters (sets of density-connected points).
Noise points (not assigned to any cluster).
Workflow Summary#
Choose parameters \(\epsilon\),
minPts.Visit each point, check density condition.
Grow clusters from core points.
Assign border points.
Mark leftover points as noise.
import numpy as np
import matplotlib.pyplot as plt
from sklearn.datasets import make_moons
from sklearn.cluster import DBSCAN
# Generate toy dataset
X, _ = make_moons(n_samples=150, noise=0.05, random_state=42)
# Parameters
eps = 0.25
minPts = 5
# Run DBSCAN for full result
db = DBSCAN(eps=eps, min_samples=minPts).fit(X)
labels = db.labels_
# Step-by-step visualization of workflow
fig, axs = plt.subplots(1, 4, figsize=(18, 4))
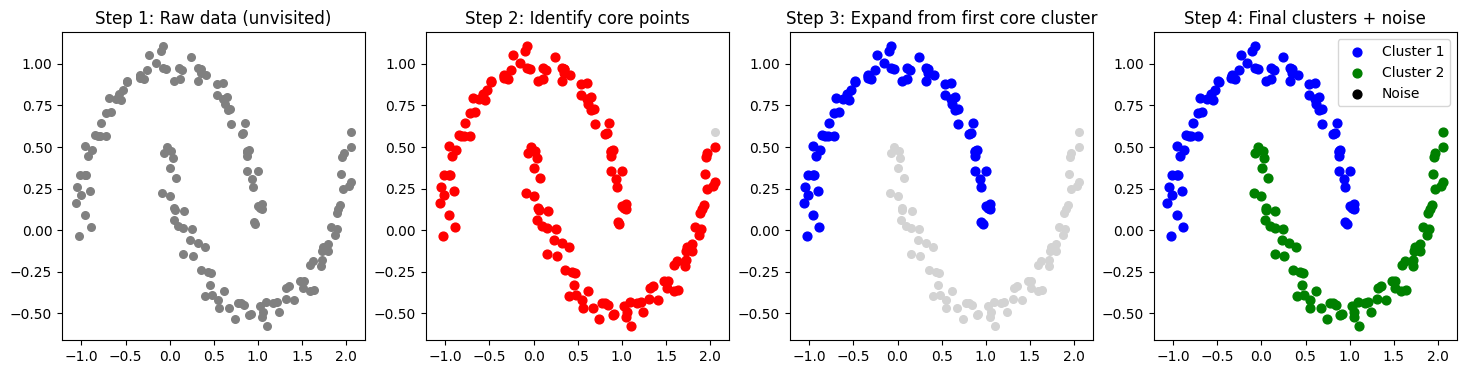
# Step 1: Raw data
axs[0].scatter(X[:, 0], X[:, 1], c="gray", s=30)
axs[0].set_title("Step 1: Raw data (unvisited)")
# Step 2: Core point identification
core_samples_mask = np.zeros_like(labels, dtype=bool)
core_samples_mask[db.core_sample_indices_] = True
axs[1].scatter(X[:, 0], X[:, 1], c="lightgray", s=30)
axs[1].scatter(X[core_samples_mask, 0], X[core_samples_mask, 1], c="red", s=40)
axs[1].set_title("Step 2: Identify core points")
# Step 3: Cluster expansion (show clusters growing)
axs[2].scatter(X[:, 0], X[:, 1], c="lightgray", s=30)
axs[2].scatter(X[labels == 0, 0], X[labels == 0, 1], c="blue", s=40)
axs[2].set_title("Step 3: Expand from first core cluster")
# Step 4: Final clusters + noise
axs[3].scatter(X[:, 0], X[:, 1], c="lightgray", s=30)
axs[3].scatter(X[labels == 0, 0], X[labels == 0, 1], c="blue", s=40, label="Cluster 1")
axs[3].scatter(X[labels == 1, 0], X[labels == 1, 1], c="green", s=40, label="Cluster 2")
axs[3].scatter(X[labels == -1, 0], X[labels == -1, 1], c="black", s=40, label="Noise")
axs[3].legend()
axs[3].set_title("Step 4: Final clusters + noise")
plt.show()